TCRβ diversity has been theoretically estimated in the range of 10 12–10 15, whereas the experimentally estimated TCRβ diversity of an adult has ranged from 10 5 to 10 8 by traditional Sanger sequencing or by next-generation sequencing. As a result of its immense size and complexity, the precise characteristics of the TCR repertoire have not yet been fully determined. The abundance of particular TCR clones reveals the immunologic history, immune status relative to pathogens, and environment of the host. The enormous diversity of TCRs ensures successful recognition of all potential antigens and serves as a measure of immune-system competency. TCR repertoire diversity can be characterized by the estimated number of unique TCRs in the total repertoire, a concept known as species richness. Conventional TCRs consist of α and β chains that are generated by random recombination among numerous gene segments, creating TCRs of vastly different specificity, collectively called the TCR repertoire.
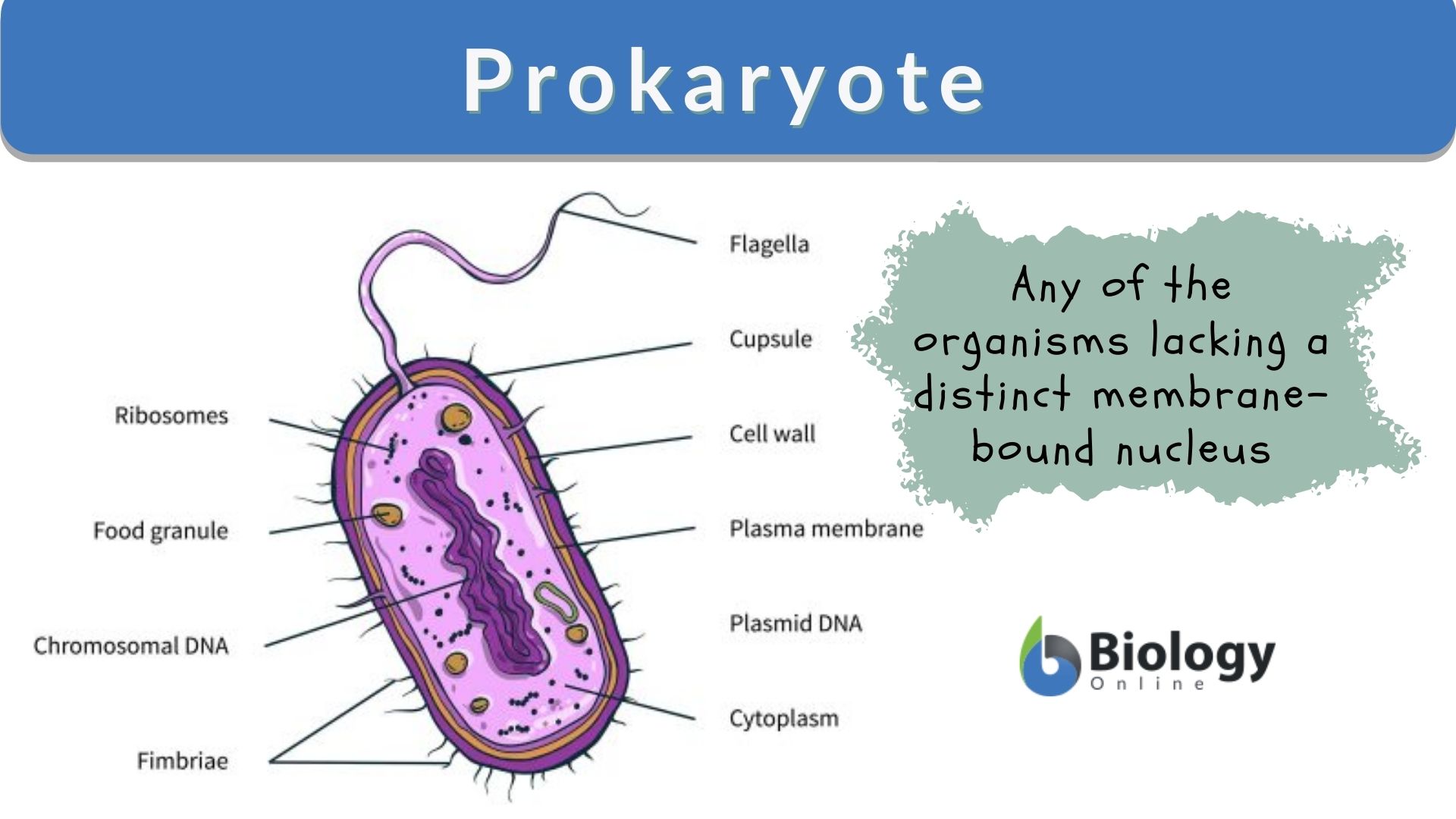
TCRs provide the specificity of the interaction between T cell and antigen–MHC complex. hematopoietic stem cell transplantation.Together, our findings reveal the distinct features of the TCRβ repertoire between CD4 + and CD8 + T cells and could potentially be used to evaluate the competency of T cell immunity. Finally, we identified 5–12% of the unique TCRβs that share an identical CDR3 with different variable genes. Further analysis showed that CD4 + and CD8 + T cells exhibited distinct preferences for certain amino acids in the CDR3, and this was confirmed further by a support vector machine classifier, suggesting that there are distinct and discernible differences between TCRβ CDR3 in CD4 + and CD8 + T cells. Furthermore, there was little overlap in TCRβ sequences between CD4 + (0.3%) and CD8 + (1.3%) T cells. We found that TCRβ richness of CD4 + T cells ranges from 1.2 to 9.8 × 10 4 and is approximately 5 times greater, on average, than that of CD8 + T cells in each study subject. Here, we report a comparative analysis of the TCRβ repertoires of CD4 + and CD8 + T cells by use of a 5′ rapid amplification of cDNA ends–PCR–sequencing method.

However, it is currently unclear to what extent the TCR repertoire of CD4 + and CD8 + T cells is different. Two major types of T cells, CD4 + and CD8 +, that use the same genetic elements and process to generate a functional TCR differ in their recognition of peptide bound to MHC class II and I, respectively. The TCR repertoire serves as a reservoir of TCRs for recognizing all potential pathogens.
